Skeletal myopathy in a family with lamin A/C cardiac disease
Introduction
Over the past 5 decades, cardiologists at a large referral center have evaluated a large kindred who present with cardiac signs and symptoms due to cardiac conduction system and myocardial disease. Affected family members share a unique autosomal dominant frameshift mutation in LMNA (chromosome 1q21.2) (1-4). The progressive cardiovascular disease course in affected members of this family has been well defined. The initial manifestations are atrial arrhythmia, which usually occur in the second and third decades of life, and subsequent PR interval prolongation on electrocardiography. Progressive atrioventricular conduction block and a broad spectrum of ventricular dysfunction occur in the fourth to fifth decades. Sudden cardiac death (SCD) has occurred in 31 confirmed genotype-affected patients, including 11 members with functioning pacemaker at the time of death (1,3,4). The incidence of SCD in this family has subsequently been attenuated with implantable cardiac defibrillator implantation.
LMNA encodes lamin A/C, and other mutations are associated with autosomal dominant Emery-Dreifus muscular dystrophy (EDMD2), limb-girdle muscular dystrophy and familial partial lipodystrophy; patients with these disorders may develop cardiomyopathy later in life (5). For instance, patients with EDMD2 with distinct LMNA mutation typically develop skeletal myopathy with contractures in childhood, and when they occur, cardiac arrhythmias and heart block develop later in life (6,7).
Skeletal muscle magnetic resonance imaging (MRI) was shown by Mercuri et al. to identify abnormalities of the calf muscles unique to EDMD2, predominantly the medial head of the gastrocnemius muscle (6). They demonstrated that skeletal muscle MRI accurately differentiated EDMD2 from non-LMNA skeletal myopathies (6). Since cardiac manifestations have been the predominant phenotype in this LMNA family, skeletal myopathy has not been previously described. We hypothesized that skeletal muscle imaging in genotype-affected laminopathy patients would identify involvement in a pattern similar to that seen in EDMD2.
Methods
Study population
Members of a family sharing an identical frameshift mutation in lamin A/C who had been genotyped as part of a longitudinal study were prospectively enrolled. Of 36 genotype-positive individuals, 21 consented to participate in this protocol. Those not participating declined primarily due to distance of current residence from the study site. Control subjects were healthy volunteers with no history of cardiomyopathy or skeletal muscle disorder. Written informed consent was obtained from all subjects participating in this Institutional Review Board-approved protocol; all exams and procedures were performed in accordance with institutional guidelines. Patients without pacemakers, defibrillators, severe claustrophobia, ferromagnetic foreign body, or aneurysm clip underwent calf muscle MRI. This included 11 LMNA patients and 17 age-matched control subjects. The remaining ten LMNA patients had pacemakers due to advanced cardiac conduction disease and instead underwent calf muscle imaging using computed tomography (CT).
Image acquisition
MRI examinations were performed using a conventional 1.5 Tesla clinical scanner (CVi, GE Healthcare). Acquisition technique consisted of standard T1-weighted fast spin echo (T1W-FSE) imaging, T1W-FSE with fat suppression, and delayed post-gadolinium enhancement imaging using an inversion recovery-prepared gradient echo sequence initially developed to identify myocardial fibrosis (8). All images were acquired in the axial plane i.e., cross-sections relative to the long axis of the leg. In LMNA patients with MR contraindication undergoing CT, non-contrast CT images were acquired in the cross-sections through the mid-calf. CT acquisitions were all performed on the identical multi-detector scanner (Somatom64, Siemens) using non-ECG gated helical acquisition spanning bilateral calves. Radiation exposure was not measured but can be estimated to be <5 mSv based on the low tube voltage (70 kVp) used and minimum tube current.
Data analysis
Presence or absence of skeletal muscle abnormality was determined both qualitatively and quantitatively. The qualitative assessment was based on identification of macroscopic fat and atrophic changes in the muscle belly by two independent investigators (SVR and SG), who were blinded to subject history (i.e., patient vs. control), with the plan of resolving any difference in opinion through consensus. There was complete correlation between the two investigators without any other technical reason to account for the striking imaging abnormalities that were noted in the skeletal muscles. Quantification of skeletal muscle MRI and CT data was done by off-line measurement of signal intensity in regions of interest (ROIs) (Siemens AG, Leonardo VD10B Syngo). ROIs encompassed the medial head of the gastrocnemius muscle and unaffected anterior compartment muscle for each image acquired using each of the three imaging techniques. Signal intensity ratios, gastrocnemius to unaffected muscle, were calculated for each of the three MRI techniques [T1W, T1W-FS, delayed enhancement (DE)]. ROIs similarly drawn on CT images yielded mean Hounsfield Unit values within gastrocnemius muscle and control muscle. A nonparametric exact Wilcoxon Rank Sum test was used to compare data between patients and controls with respect to each continuous variable. A value of P<0.05 was regarded as statistically significant. Comparison of frequencies was done using Fisher’s exact test.
Results
Basic demographics of the study population are given in Tables 1-3. Consistent with the progressive nature of the cardiac conduction disease in this mutation, the LMNA patients undergoing CT due to presence of pacemakers were significantly older compared to the LMNA patients able to undergo MRI due to lack of pacemaker (mean age 57 vs. 33 years, P<0.0001).
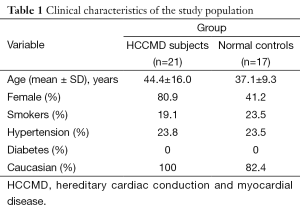
Full table

Full table

Full table
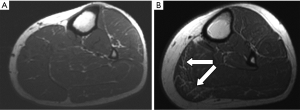
Qualitative assessment of calf muscle imaging ranged from normal in 8/11 LMNA patients to visibly abnormal in 3/11 LMNA patients (Figure 1), notably the three oldest patients in the MRI group. Increased signal intensity in the region of the gastrocnemius muscle on T1W imaging was reduced with fat suppression and enhanced on delayed post-gadolinium enhancement imaging consistent with fibrofatty replacement of normal skeletal muscle (Figure 2). Calf muscle appeared strikingly abnormal in 9/10 LMNA patients undergoing CT (Figure 3). No control subjects’ calf muscle images appeared abnormal. Fisher’s exact test showed a significant difference in frequency of qualitatively apparent skeletal muscle abnormality in patients vs. controls (P<0.05).
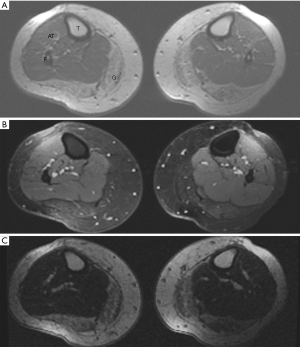
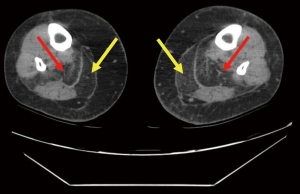
Quantitative analysis T1W images revealed similar ratio of gastrocnemius muscle to unaffected muscle signal intensity in LMNA MRI patients vs. MRI control subjects (1.16±0.3 vs. 1.29±0.6, respectively, P= NS). For the fat suppressed images, the ratios were 1.1±0.84 in LMNA MRI patients and 1.3±0.3 in controls (P= NS). Lastly the ratios for DE calf images in LMNA MRI patients vs. MRI controls were 1.4±0.85 and 1.13±0.4, respectively (P= NS). In LMNA patients undergoing CT, the mean Hounsfield unit (HU) value in the gastrocnemius muscle was significantly lower than that measured in the unaffected anterior tibialis (–18.6±38 vs. 59.2±8.0 HU, P<0.001).
Discussion
We found evidence of skeletal myopathy in genotype-affected members of a family with heritable cardiac conduction and myocardial disease due to mutation in LMNA. These findings were previously undetected as their signs and symptoms such as palpitations, syncope, heart failure, and SCD have focused evaluation and management on their cardiac disease. This investigation was prompted by appreciation that both the primarily cardiac disease in this family and the primarily skeletal muscle disease in EDMD2 result from mutations within LMNA. Unlike our family, patients with EDMD2 typically develop advanced skeletal myopathy well before the onset of cardiomyopathy (6,7).
Lamin A/C is an intermediate filament cytoskeletal protein that maintains the structural integrity of the nuclear envelope and plays a critical role in organizing chromatin within the nucleus (9). As described in EDMD2 patients (6), our LMNA patients with skeletal muscle abnormalities demonstrated involvement primarily of the gastrocnemius muscle. Mechanisms that produce skeletal muscle involvement are unknown in both populations, and can be attributed to variable gene expression (10). The mechanical stress model could explain differential muscle via greater weight-bearing on the posterior compartment calf muscles (11). As LMNA may regulate transcriptional response to mechanical stress, it could be that the greater stress on the calf muscles accelerates myocyte loss and subsequent fibrofatty replacement. Demonstration of higher prevalence of skeletal muscle abnormality in older LMNA patients undergoing CT compared to younger LMNA patients who could safely undergo MRI is consistent with the notion of stress-related muscle loss that increases over time. The parallel between more advanced skeletal muscle changes and more advanced cardiac disease is also consistent with a physical stress-related loss of cardiac myocytes known to result in cardiac fibrosis (4) as well as loss of skeletal myocytes. Cardiac magnetic resonance allows us to detect myocardial fibrosis in vivo in these patients (12); image guidance may facilitate future discovery of molecular mechanisms that influence onset and severity of myocardial vs. skeletal muscle fibrosis.
Our study has some limitations, most importantly related to small sample size. We only evaluated consenting members of a single family who were affected by a unique frameshift mutation in the LMNA gene (chromosome 1q21.2). While this may lead to potential selection bias, we feel that the analysis of results extends beyond the affected family members. This is because correlation between skeletal and myocardial disorders is well recognized in similar patient populations, which are also affected by mutations of the LMNA gene. While, skeletal myopathy has been known in primary LMNA-related dystrophies, it has not been consistently recognized in patient cohorts where cardiomyopathy dominates the presentation (e.g., with arrhythmias and heart failure). LMNA encodes lamin A/C, mutations in which are associated with autosomal dominant EDMD2, limb-girdle muscular dystrophy and familial partial lipodystrophy. In these disorders, patients develop skeletal myopathy in childhood and may develop cardiomyopathy later in life. In our patients, findings of skeletal myopathy were previously unrecognized as they presented with cardiac signs and symptoms, thus prompting a focused evaluation and management of their cardiac disease. The identification of skeletal muscle changes in these patients with known cardiomyopathy may alert the clinician to consider skeletal myopathy as the cause of patient-reported symptoms such as weakness and exertional fatigue, which may previously have been attributed solely to cardiac disease. We recognize that our findings were in a single family with a shared mutation, and applicability to other LMNA cohorts cannot be assumed without further investigations. Nonetheless, given the overlap in cardiac and skeletal muscle involvement in other diseases known to affect both cardiac and skeletal muscle with a range of mutations affecting the same gene (e.g., Duchenne muscular dystrophy, Becker muscular dystrophy, etc.), we anticipate that our findings will prompt identification of skeletal myopathy in other cardiomyopathy-predominant LMNA cohorts.
As this was a hypothesis-generating exploration of skeletal muscle in a cohort being primarily evaluated for cardiac disease, additional investigations were not performed in this group including correlation of non-invasive imaging with EMG or other neurological measures such quantitative strength testing. CPK levels which are known to be elevated in LMNA cardiomyopathy were not routinely measured in our patients at the time of imaging of skeletal muscles. Future studies may focus on correlation of imaging data with levels of muscle enzymes in these subsets of patients. In other conditions producing myocardial and skeletal muscle disease, measurement of CPK has largely been abandoned due to lack of specificity and diagnostic value. Additionally we did not include NMR spectroscopy to assess for metabolic dyscrasias in the affected tissues; it may be that fibrofatty replacement follows earlier metabolic abnormalities triggered by abnormal LMNA- related processes that can be detected with spectroscopic measurements of energy utilization (13). We do not have data on unaffected mutation carriers. This was beyond the scope of the study but an important path to consider for future investigations.
Conclusions
In summary, we found previously undetected skeletal myopathy in genotype-affected family members with cardiac conduction and myocardial disease due to LMNA mutation. The extent of skeletal muscle involvement was more pronounced in older LMNA family members with advanced cardiac disease. Additional investigations are warranted to define the factors that dictate cardiomyopathy and skeletal myopathy phenotypic variations among the laminopathies.
Acknowledgements
This work was supported in part by a Davis Medical Research Grant (SVR). We are grateful to the patients and their families for their participation in this work. We dedicate this work to the memory and continued inspiration of Charles F. Wooley, MD.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
Ethical Statement: All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Informed consent was obtained from all individual participants included in the study.
References
- Graber HL, Unverferth DV, Baker PB, et al. Evolution of a hereditary cardiac conduction and muscle disorder: a study involving a family with six generations affected. Circulation 1986;74:21-35. [Crossref] [PubMed]
- Sparks EA, Graber HL, Boudoulas H, et al. Atrial myopathy and atrial fibrillation: phenotypes in heritable cardiac conduction and myocardial disease. Eur Heart J Suppl 2000;2:K79-K90.
- Kass S, MacRae C, Graber HL, et al. A gene defect that causes conduction system disease and dilated cardiomyopathy maps to chromosome 1p1-1q1. Nat Genet 1994;7:546-51. [Crossref] [PubMed]
- Nelson SD, Sparks EA, Graber HL, et al. Clinical characteristics of sudden death victims in heritable (chromosome 1p1-1q1) conduction and myocardial disease. J Am Coll Cardiol 1998;32:1717-23. [Crossref] [PubMed]
- Charniot JC, Pascal C, Bouchier C, et al. Functional consequences of an LMNA mutation associated with a new cardiac and non-cardiac phenotype. Hum Mutat 2003;21:473-81. [Crossref] [PubMed]
- Mercuri E, Counsell S, Allsop J, et al. Selective muscle involvement on magnetic resonance imaging in autosomal dominant Emery-Dreifuss muscular dystrophy. Neuropediatrics 2002;33:10-4. [Crossref] [PubMed]
- Morris GE, Manilal S. Heart to heart: from nuclear proteins to Emery-Dreifuss muscular dystrophy. Hum Mol Genet 1999;8:1847-51. [Crossref] [PubMed]
- Kim RJ, Shah DJ, Judd RM. How we perform delayed enhancement imaging. J Cardiovasc Magn Reson 2003;5:505-14. [Crossref] [PubMed]
- Ostlund C, Worman HJ. Nuclear envelope proteins and neuromuscular diseases. Muscle Nerve 2003;27:393-406. [Crossref] [PubMed]
- Kang PB, Kho AT, Sanoudou D, et al. Variations in gene expression among different types of human skeletal muscle. Muscle Nerve 2005;32:483-91. [Crossref] [PubMed]
- Worman HJ, Courvalin JC. How do mutations in lamins A and C cause disease? J Clin Invest 2004;113:349-51. [Crossref] [PubMed]
- Raman SV, Sparks EA, Baker PM, et al. Mid-myocardial fibrosis by cardiac magnetic resonance in patients with lamin A/C cardiomyopathy: possible substrate for diastolic dysfunction. J Cardiovasc Magn Reson 2007;9:907-13. [Crossref] [PubMed]
- Kemp GJ, Taylor DJ, Dunn JF, et al. Cellular energetics of dystrophic muscle. J Neurol Sci 1993;116:201-6. [Crossref] [PubMed]


