
Implications of gut microbiome on coronary artery disease
Introduction
Coronary artery disease (CAD) is currently the number one killer of human health worldwide, including a series of heart and vascular dysfunction disorders, the most important of which is atherosclerosis (AS) (1). With the increasing trend of population aging, the incidence of AS has increased dramatically. According to the World Health Organization, CAD will be the world’s leading cause of death by 2020. Currently, most of the biomarkers for screening cardiovascular disease (CVD) are based on pathways such as inflammation and cholesterol biosynthesis. Furthermore, lipid-rich diets have a clear correlation with the progression of CVDs, so the main treatment direction is to reduce the intake or synthesizes of cholesterol, and reduces the intake of trans fatty acids, saturated fatty acids and triacylglycerols in the diet. However, there are reports that patients with active statin therapy, even if low-density lipoprotein cholesterol (LDL-C) has reached the standard, which below 70 mg/dL, the residual cardiovascular risk is still present (2). Consequently, there is an increasing need to identify novel therapeutic targets and develop new anti-atherosclerotic drugs.
With the continuous development of gene sequencing, metabolomics, proteomics, and bioinformatics, people’s research on the gut microbiome has been deepened, and the mystery of gut microbiome has been gradually unveiled. A large number of studies have shown that gut microbiome dysbiosis is closely related to CVDs and its risk factors such as diabetes mellitus (3), obesity (4), and AS (5) over the past decade. The role of the gut microbiome in the occurrence and development of CAD has drawn increasing attention, as well as gradually been considered as a potential diagnostic and pharmacological target for the prevention and treatment of CAD (6).
Here we provide a brief overview of gut microbiome composition in human physiology and present the recently reported evidence establishing the association between alterations in gut microbiome and metabolites and CAD. Finally, we will highlight the alteration of the gut microbiome for prognostic and preventive purposes in CAD.
Overview of the gut microbiome
The human intestine contains >1014 of the gut microbiome, which including bacteria, archaea, yeast, viruses that are dependent on the human gut and help the host complete multiple physiological and biochemical functions accompanied by their metabolites (7). The term gut microbiota refers to all the microorganisms colonized in the intestinal tract, and gut microbiome collectively means the genes and genomes of the microbiota as well as their by-product and the host environment (8). The composition of a healthy individual’s gut microbiome can vary remarkably from another’s (9), however, this composition is relatively stable over time (10). The gut microbiome is primarily composed of species within the Bacteroidetes, Firmicutes, Actinobacteria, and Proteobacteria, with over 90% of the total bacterial species in the healthy bacterial community are Bacteroidetes and Firmicutes (11). The relative abundance of species varies among individuals due to a variety of genetic and environmental factors, including dietary habits, intestinal infection and the administration of antibiotic drugs.
The balance of gut microbiome composition helps to protect the intestinal mucosal barrier, facilitates nutrition intake and metabolism regulations, assists immune tissue maturity, and prevents pathogenic microorganisms from entering the systemic circulation (12). Alterations in the gut microbiome may cause gut microbiome dysbiosis, damage the integrity of the intestinal mucosal barrier, and increase the level of systemic inflammation, thereby affecting the health of the host. The gut microbiome produces a large number of metabolites through food absorption and digestion, some bioactive metabolites can act on distant target organs in a manner similar to human endocrine organs (13). For example, lipopolysaccharides (LPS) and peptidoglycans produced by intestinal flora metabolism, interact with host mucosal surface cells through pattern recognition receptors, recognize molecular patterns related to pathogens, as well as stimulate the host's immune response (14).
Relationship between gut microbial composition and CAD
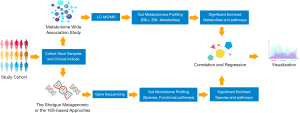
Previously, microbial research was conducted in a traditional culture-based approach, which could provide us with direct information regarding living conditions and metabolism (15). However, this method is only available to identify those culturable bacterial strains, which comprise just a slight part of the human gut microbial community. With advances in next-generation sequencing, new strategies including 16S ribosomal RNA (rRNA) sequencing and metagenomics have brought us new insights to explore a more complex microorganism system (16,17). 16S rRNA sequencing depends on the polymerase chain reaction (PCR) amplification of a specific region in the 16S gene for the taxonomic identification of the bacteria community (18). Although robust and well-characterized, the limitations of this method include that taxa are assigned on the results of the sequence of a single region of the bacterial genome, different region sequencing can generate different results. Moreover, the sequencing of the regions of variability is generally inadequate to provide species-level resolution (6). The metagenomic shotgun sequencing analyses broad genomic regions and more complex downstream data with potential functional profiles via computational bioinformatics (19). It enables a more accurate taxonomic identification at the species level, thus yielding a theoretically comprehensive and insightful description of the bacterial community. The flowchart of the contemporary strategies for gut microbiome and metabolomics analysis in human cohorts were shown in Figure 1 (20). According to the current studies, almost every CVD has shown associations with the alterations in gut microbial composition (Table 1).
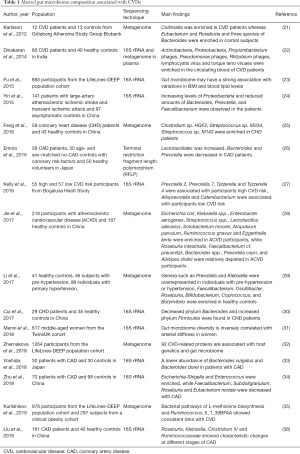
Full table
Multiple studies have been carried out to identify microbial strains and functions associated with CAD. Kelly et al. conducted 16S rRNA sequencing on microbial DNA from stools of participants with different lifetime burdens of CVD risk factors, and the results showed that Prevotella 2, Prevotella 7, Tyzzerella and Tyzzerella 4 were enriched in participants high CVD risk, Alloprevotella and Catenibacterium were enriched participants low CVD risk (27). Jie et al. performed a metagenomic shotgun-sequencing on stool samples from 218 patients with atherosclerotic CVDs and 187 healthy controls and identified an increased abundance of Enterobacteriaceae and Streptococcus spp. in atherosclerotic CVDs (28). Another study conducted in Japan, which involved 30 CAD patients and 30 non-CAD controls, revealed a negatively correlated abundance of Bacteroides vulgatus and Bacteroides dorei in CAD patients by fecal 16S rRNA sequencing (33). The reasons why discrepancies on microbial signatures of different studies are due to (I) the different study population toward a region-specific genetic background and dietary habits, and both are known to affect metabolism and the gut microbiome (27); (II) difference in sequencing strategies. 16S rRNA sequencing has low phylogenetic power at the species level and poor discriminatory ability for some genera (33); (III) the intrinsic flaw of taxon-based analysis, which overlooks the variations of the bacterial strains belonging to the same taxon (36); (IV) the association analysis based on a cross-sectional design has underlying causality and the mechanism remains unclear (35). Several studies have reported that genetics and microbiome background can exert independent additive effects on CVD-related phenotypes including BMI, blood lipid levels and so on, which may limit the power of randomization analysis to illustrate underlying causality (23,37). Nevertheless, mounting evidence has shown the key members of the gut microbiome might be potential candidates in the development, as well as in animal models. Yoshida et al. treated six-week-old female ApoE−/− mice, which were on C57BL/6 background, with live Bacteroides vulgatus and Bacteroides dorei by oral gavage and found that compared with control mice, mice gavaged with live Bacteroides showed significant atherosclerotic lesion size along with a reduction in macrophage and CD4+ T cell accumulation as well as the mRNA expression of proatherogenic immune cell makers chemokine receptors (33). Li’s team reported recipient germ-free mice inoculated with stool samples from hypertensive donors showed a tendency for higher blood pressure, indicating that high blood pressure is transferrable by fecal transplant (29). These findings may provide new insights for revealing novel potential aetiologies for CAD, understanding the role of the gut microbiome in CAD, and modulating gut microbiome as a therapeutic target.
Gut microbial metabolites as regulators in CAD
Trimethylamine-N-oxide (TMAO)
The gut microbiome produces trimethylamine (TMA) by ingesting choline (such as choline, phosphatidylcholine, and L-carnitine) in food. TMA enters the liver through the enterohepatic circulation, where oxidized by flavin-dependent monooxygenase isoforms 1 and 3 (FMO1 and FMO3) to form TMAO, and then enters the systemic circulation (38). TMAO has been considered to be a promising promotor of AS, when choline-rich food is fed to mice with normal gut microbiome, TMAO levels in the circulation increase, causing foam cells to aggregate and promoting atheromatous plaque formation (39). Inhibition of gut microbiota-independent TMAO production has been shown a novel therapeutic strategy for the treatment of CAD (40). Ma’s team reported that TMAO promotes monocyte adherence by up-regulating the level of vascular cell adhesion molecule-1 (VCAM-1), and activates the protein kinase C (PKC) and nuclear factor κB (NF-κB) pathways, which disrupts the function of endothelial cells and accelerates the early pathological process of AS (41). Elevated TMAO levels can affect mitochondrial repair and myocardial metabolism, as well as significantly increase the risk and severity of acute myocardial infarction (42). A study involving over 4,000 participants demonstrated that plasma TMAO level can predict the risk of a 3-year thrombosis incident (heart attack, stroke). Moreover, TMAO can also augment the intracellular release of Ca2+ through adenosine diphosphate (ADP), thrombin and collagen, increase the responsiveness of platelets, and then promote thrombosis, which may directly stimulate the occurrence of myocardial infarction (41). Additionally, in the complicated carnitine biosynthesis pathway, precursors of carnitine, γ-butyrobetaine (γBB), and trimethyllysine (TML) were also adequately investigated as potential substrates for TMA/TMAO production (43,44).
Short-chain fatty acids (SCFAs)
Organic fatty acids with 1 to 6 carbon atoms are called SCFAs, mainly including acetic acid, propionic acid, isobutyric acid, butyric acid, isovaleric acid, and valeric acid. The gut microbiome is involved in the metabolic process of dietary fiber and resistant starch to produce SCFAs. A large number of intestinal bacteria can produce acetic acid, a few members of the families Veillonellaceae and Lachnospiraceae are propionic acid-producing bacteria, Coprococcus, Faecalibacterium, Eubacterium, and Roseburia are butyrate-producing bacteria (45). The human gastrointestinal tract generally produces SCFAs of about 50 to 100 mmol per day, most of which are derived from undigested carbohydrates fermented by intestinal bacteria, and a small part are derived from daily diet and protein metabolism. SCFAs in the intestine mainly exist in ionic form, which can enter the circulatory system through the hepatic and enterohepatic circulation (46). Although SCFAs account for approximately 2% to 10% of energy consumption in humans, they also serve as signaling molecules to modulate autonomic systems and blood pressure, as well as inflammatory responses and other cellular functions. SCFAs exert a broad range of physiological functions, including histone deacetylases inhibition, chemotaxis and phagocytosis modulation, reactive oxygen species induction, and immunoregulation in monocytes and macrophages (47). Studies show that SCFA have multiple receptors, including G-protein coupled receptors 41 (GPR41), GPR43, GPR109A, and olfactory receptor 78 (Olfr78) (48). SCFAs (especially propionate and butyrate) can affect the body’s immune and metabolic functions by regulating these receptor signaling pathways, inhibiting NF-κB and tumor necrosis factor (TNF) signaling pathways, leading to a decreased expression of VCAM-1 and intercellular adhesion molecule-1 (ICAM-1), thereby inhibiting the occurrence of AS (49). However, direct demonstration of such effects in human CAD remains limited.
Bile acids (BAs)
BAs are amphiphilic molecules synthesized by cholesterol in the liver. This process is an important way to eliminate cholesterol in the body. The BAs stored in the gallbladder are secreted into the intestine, which promotes the emulsification of dietary fat and helps the intestine to absorb lipid nutrients and fat-soluble vitamins. As a signal molecule, BAs mainly bind to two receptors, the farnesoid X receptor (FXR) and the G protein-coupled receptor (Takeda-G-protein receptor 5, TGR5) (50,51). FXR is a bile acid-activated transcription factor, mainly expressed in the liver, kidney, and intestines, can be expressed by its downstream target genes, small heterodimer partner (SHP) and fibroblast growth factor (FGF) 15 or FGF19 (mouse FGF15, rat/human FGF19) regulates transintestinal cholesterol excretion (52). FXR activation increases fecal cholesterol excretion and reverse transport of cholesterol by macrophages, and at the same time inhibits intestinal absorption of cholesterol, lowers cholesterol and triglycerides in plasma, thus becoming one of the therapeutic targets for dyslipidemia (53). Studies have confirmed that FXR and TGR5 dual bile acid receptor agonist INT-767 can inhibit NF-κB pathway, which significantly reduced circulating lipids such as LDL-C and the expression of aortic inflammatory cytokines and chemokines to exert anti-inflammatory effects (54).
The role of microbial dysbiosis in atherosclerotic process and plaque stability
Possible mechanisms of gut microbial composition and metabolites participate in the progression of CAD
The chronic atherosclerotic process initiates with accumulation of low-density lipoprotein and fibrous substances, coupling with inflammatory responses in all phases. The gut microbiome is closely related to the formation and development of AS, it will affect the stability of atherosclerotic plaques. Karlsson et al. performed a metagenomic analysis which compared unstable plaques with stable plaques and found that patients with unstable plaques have a lower abundance of Roseburia in their fecal samples (21). Koren et al. reported that the bacterial DNA in arterial plaques is similar to that of the gut microbiome, and the bacteria in these plaques may be related to plaque stability (55). Additionally, a recent study has reported that the relative abundance of phylum Zygomycota and Mucor racemosus are inversely related to the carotid intima-media thickness and to the occurrence of subclinical AS (56).
Furthermore, the range of intestinal microbiota metabolites extends far beyond the local environment of the intestine, thus affecting the physiological functions of remote organs such as the heart (57). Gut microbiome dysbiosis leads to decreased expression of tight junction protein, increased permeability of intestinal mucosa, and LPS components on the outer membrane of Gram-negative bacteria enter the blood circulation (58). LPS induces the expression of chemokines and cell adhesion molecules, promotes the formation of foam cells in the adhesion of monocytes to the endothelial cells, and starts the process of AS (59). In addition, LPS can also directly bind to toll-like receptor 4 (TLR4) on the surface of immune cells, inhibit liver X receptor (LXR), and reduce adenosine triphosphate (ATP) binding cassette transporters (ABCA1 and ABCG1). ABCA1 and ABCG1 are integrated membrane proteins that use ATP as an energy source to promote the outflow of free cholesterol and phospholipids in cells and play an important role in reverse cholesterol transport (RCT). Secondly, LPS can also indirectly trigger the release of pro-inflammatory factors, such as TNF-α, interleukin (IL)-1β, inhibit the expression of cholesterol transporters, and then promote the formation of foam cells (56). Besides, butyrate and other SCFA produced by gut microbiota can directly feed colonocytes which contribute to preventing inflammation and gut leakage (60). Jie et al. reported a reduced abundance of Roseburia Intestinalis and Faecalibacterium prausnitzii, which are known producers of SCFA butyrate, in patients with atherosclerotic CVD (28). In accordance with Jie’s study, another two studies also shown that butyrate producer Roseburia and Eubacterium are significantly depleted with CAD (21,34). Along this line of consideration, the elevated gut permeability accounts for the increased intestinal microbiota translocation with higher LPS level into the systemic circulation, which has been reported to associated with adverse outcomes in post myocardial infarction patients (61). Accumulating studies have confirmed TMAO to interfere with the process of CAD, additionally, plasma TML level, alone and in combination with TMAO are reported to be significantly associated with both near- and long-term cardiovascular events, independent of troponin T levels (62).
Overall, the gut microbiome can affect the occurrence and development of AS in a variety of ways, mainly including (I) gut microbiome dysbiosis may lead to inflammatory response, aggravate the development of atherosclerotic plaque or cause plaque rupture; (II) gut microbiome affects the development of atherosclerotic plaques by regulating host cholesterol and lipid metabolism; (III) metabolites of the gut microbiome may have a beneficial or harmful effect on the occurrence of AS.
Oral microbial disorders and periodontal disease are also related to CAD
Periodontal disease is a common bacterial infection that causes chronic inflammation and host immune response to accelerate AS (63). A study reported that bacterial DNA, which can also be detected in the oral cavity, exists in the plaques of CAD patients (64). Besides, Renvert et al. assayed subgingival pathogens in patients admitted with ACS and discovered a higher oral bacterial load in ACS patients compared with matched controls, the mostly increased species including Porphyromonas gingivalis, Tannerella forsythensis, Treponema denticola, Streptococcus intermedius, and Streptococcus sanguis (65). A meta-analysis included 15 clinical studies involving 17,330 participants confirmed the correlation between periodontal disease and carotid AS (66). Another meta-analysis included 17 case-control studies involving 3,456 patients with myocardial infarction and 3,875 non-myocardial infarction controls, showing that myocardial infarction patients had worse periodontitis and oral hygiene than non-myocardial infarction patients (67). Further large-scale, well-designed clinical studies focusing on the causal relationship between CAD, ACS in particular, and periodontal disease should be conducted in the future.
Novel potential therapeutic interventions targeting the gut microbiome
Targeting gut microbiome and its metabolites are becoming a novel and attractive field in the treatment of CAD. In a rat model of myocardial infarction, the use of broad-spectrum antibiotic drugs has affected the composition of the gut microbiome, and the infarct size has been reduced by affecting the levels of aromatic amino acid catabolites and leptin (54). Additionally, Adding Lactobacillus plantarum 229v to food as a probiotic supplement can significantly reduce circulating leptin levels and myocardial infarct size in rats (68). Patients with myocardial infarction were detected bacterial DNA (Lactobacillus, Bacteroides, and Streptococcus) in the blood, demonstrating the translocation of gut-originated bacteria into the systemic circulation, which consequently predisposes patients to cardiovascular events. This study also revealed that the administration of antibiotics to reduce the translocation gut microbial translocation post-myocardial infarction can alleviate the systemic inflammation and myocardial injury (61). However, improper use of antibiotics can kill beneficial GM, we should pay attention to the side effects of antibiotics and their clinical effects.
Although harmful intestinal bacteria can promote the occurrence of AS, there are also a large number of beneficial bacteria in the intestinal microorganisms, which can have a good prevention and treatment effect on the occurrence and development of AS. Chen et al. treated ApoE−/− mice with Lactobacillus acidophilus ATCC 4356 for 12 weeks and found that compared with model control animals, the body weight and serum lipid profile of the treated mice did not change, but showed decreased atherosclerotic lesion size. In addition, Lactobacillus acidophilus ATCC 4356 dose-dependently reduced the levels of malondialdehyde (MDA), oxidized low-density lipoprotein (ox-LDL), and TNF-α in the serum profile of treated mice, increased serum interleukin-10 (IL-10) levels and superoxide dismutase (SOD) activity (69). Akkermansia muciniphila, a mucin-degrading bacterium with beneficial effects on obesity-related metabolic disorders, attenuates atherosclerotic lesions by ameliorating metabolic endotoxemia-induced inflammation through the restoration of the gut barrier in ApoE−/− mice (70). Therefore, the probiotics can slow the development of atherosclerotic lesions by reducing oxidative stress and inflammatory responses in animals. Qiu et al. also found that Lactobacillus plantarum ZDY04 can reduce serum TMAO and cecum TMA levels by regulating the relative abundance of families Lachnospiraceae, Erysipelotrichaceae, Bacteroidaceae and the genus Mucispirillum in the intestinal tract of mice, thereby inhibiting AS caused by TMAO (71).
Some other drugs and food nutrients can interact with the gut microbiome to prevent and regulate the development of AS. Wang’s team reported that protocatechuic acid (PCA), a gut microbiome metabolite of cyanidin-3 to 0-β-glucoside (Cy-3-G) in ApoE−/− mice, could promote cholesterol efflux from macrophages and macrophage ABCA1 and ABCG1 expression via the regulation of miRNA-10b-ABCA1/ABCG1 signaling cascade (72). Matziouridou et al. fed male ApoE−/− mice 44% lingonberries for 8 weeks, and reported that compared with mice fed a high-fat diet, the total cholesterol and triglyceride levels and the number of atherosclerotic plaques in the lingonberries group significantly decreased. Further analysis found that lingonberries increased the relative abundance of genera Bacteroides, Parabacteroides and Clostridium, as well as upregulated the bile acid synthesis gene Cyp7a1, which might correlate with the regulation AS (73). Accordingly, some of the nutrients in the diet are likely to be more effective in preventing and treating AS after they are metabolized by the gut microbiome. In addition to dietary nutrition, many drugs can also exert anti-atherosclerotic effects by regulating the gut microbiome. Chen et al. found resveratrol (RSV), a natural phytoalexin, attenuated TMAO-induced AS in ApoE−/− mice by inhibiting commensal microbial TMA production. Moreover, RSV increased the relative abundance of the genera Lactobacillus and Bifidobacterium, which increased hepatic BAs neosynthesis through the enterohepatic FXR-FGF15 axis (74). Zhu’s team administrated berberine to mice that fed either a normal chow diet or a high-fat diet in drinking water (0.5 g/L) for 14 weeks, and the results showed that berberine may exert anti-atherosclerotic and metabolic protective effects by increasing Akkermansia spp. abundance, decreasing the intestinal expression of proinflammatory cytokines and chemokines, and enhance the restoration of gut barrier integrity (75).
Fecal microbiota transplantation (FMT) is the transfer of donor stool flora or metabolites into the colon of recipients, providing nutrition, inhibiting the growth of pathogenic bacteria, regulating the host’s immune system, and correcting intestinal microecological imbalances, thereby helping patients to rebuild the normal function of the gut microbiome. A meta-analysis speculates that microbiota transplantation may be more effective than probiotics in restoring GM, because transplanting microbiota can overcome the short-term efficacy of probiotics and cause permanent changes in the microbiota (76). Hu et al. reported FMT treatment improved myocardial injury in the experimental autoimmune myocarditis (EAM) mouse model by diminished inflammatory infiltration, accompanied by a decreased Firmicutes/Bacteroidetes ratio and a rebalanced microbiota composition (77). However, the use of FMT is currently limited because it is risky and may transfer endotoxins or other dangerous drugs into circulation, which can lead to new complications. Although FMT is a promising treatment for CVDs, further research is needed to guarantee its safety and effectiveness.
In summary, the current prevention and treatment approaches targeting gut microbiome mainly include (I) administration of antibiotics. However, the current general view is that oral broad-spectrum antibiotics have more harm than good in the treatment of gut microbiome dysbiosis because beneficial bacteria will also be inhibited, of which the metabolic processes can contribute to cardiovascular protection; (II) administration of probiotics; (III) diet intervention and small molecule metabolites or drugs; (IV) FMT. Discovering key molecules produced by gut bacterial metabolism will help to find precise molecular therapeutic pathways. Although the current research is not enough to prove that the gut microbiome is directly involved in AS, these studies have provided us with new directions and targets, as well as provided new ideas for the prevention and treatment of CAD.
Conclusive remarks and future perspective
The gut microbiome is a huge and mysterious field. The mechanism for maintaining basic life activities of the body is complex, and more and more in-depth research is needed. Accumulating evidence from experiments and clinical studies have demonstrated that the gut microbiome plays an important role in host health and disease. In addition, the identification of bacterial metabolites has opened up a variety of microbial pathways for regulating host physiological processes, making it possible to serve as a vector and potential pharmacological target for the treatment of CAD. Although gut microbiome analyses are not yet available for clinical use (78), microbial-derived biomarkers such as LPS and TMAO are underlying prognostic targets in risk stratification. Future research directions should be to clarify the mechanism of interaction between the gut microbiome and the host, describe the maturation process of the gut microbiome during host development and its impact on the early life and adult health outcomes, clarify its role in disease pathogenesis, and evaluate its feasibility for diagnosis and treatment of disease.
Moreover, the pathophysiology of CAD is a chronic, long-term process that is coupling with inflammatory responses (79). The current clinical laboratory tests or imaging examination are still focused on detecting the biomarkers of cardiac ischemia or coronary artery stenosis, and as a result do not lead to direct optimizing preventative strategies to the majority of CAD patients who would otherwise go on to suffer an ACS. Based on the lack of early warning of CAD, how to move forward to the pre-hospital level of prognosis to those who might suffer an ACS becomes a pain point for future research, the gut microbiome may catch the trend and any progress that improve identification of these individuals will reduce the burdens on public health.
Acknowledgments
Funding: This work was supported by the Chinese Academy of Medical Science (CAMS) Innovation Fund for Medical Sciences (CIFMS, 2016-12M-1-009).
Footnote
Conflicts of Interest: Both authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/cdt-20-428). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Benjamin EJ, Virani SS, Callaway CW, et al. Heart Disease and Stroke Statistics-2018 Update: A Report From the American Heart Association. Circulation 2018;137:e67-e492. [Crossref] [PubMed]
- Cholesterol Treatment Trialists’ (CTT) Collaboration, Baigent C, Blackwell L, et al. Efficacy and safety of more intensive lowering of LDL cholesterol: a meta-analysis of data from 170,000 participants in 26 randomised trials. Lancet 2010;376:1670-81. [Crossref] [PubMed]
- Yang Q, Lin SL, Kwok MK, et al. The Roles of 27 Genera of Human Gut Microbiota in Ischemic Heart Disease, Type 2 Diabetes Mellitus, and Their Risk Factors: A Mendelian Randomization Study. Am J Epidemiol 2018;187:1916-22. [Crossref] [PubMed]
- Watanabe K, Igarashi M, Li X, et al. Dietary soybean protein ameliorates high-fat diet-induced obesity by modifying the gut microbiota-dependent biotransformation of bile acids. PLoS One 2018;13:e0202083. [Crossref] [PubMed]
- Li DY, Tang WHW. Gut Microbiota and Atherosclerosis. Curr Atheroscler Rep 2017;19:39. [Crossref] [PubMed]
- Tang WHW, Bäckhed F, Landmesser U, et al. Intestinal Microbiota in Cardiovascular Health and Disease: JACC State-of-the-Art Review. J Am Coll Cardiol 2019;73:2089-105. [Crossref] [PubMed]
- Fouhy F, Ross RP, Fitzgerald GF, et al. Composition of the early intestinal microbiota: knowledge, knowledge gaps and the use of high-throughput sequencing to address these gaps. Gut Microbes 2012;3:203-20. [Crossref] [PubMed]
- Whiteside SA, Razvi H, Dave S, et al. The microbiome of the urinary tract--a role beyond infection. Nat Rev Urol 2015;12:81-90. [Crossref] [PubMed]
- Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 2012;486:207-14. [Crossref] [PubMed]
- Faith JJ, Guruge JL, Charbonneau M, et al. The long-term stability of the human gut microbiota. Science 2013;341:1237439. [Crossref] [PubMed]
- Qin J, Li R, Raes J, et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 2010;464:59-65. [Crossref] [PubMed]
- Hamilton MK, Boudry G, Lemay DG, et al. Changes in intestinal barrier function and gut microbiota in high-fat diet-fed rats are dynamic and region dependent. Am J Physiol Gastrointest Liver Physiol 2015;308:G840-G851. [Crossref] [PubMed]
- Zhu W, Gregory JC, Org E, et al. Gut Microbial Metabolite TMAO Enhances Platelet Hyperreactivity and Thrombosis Risk. Cell 2016;165:111-24. [Crossref] [PubMed]
- Schokker D, Fledderus J, Jansen R, et al. Supplementation of fructooligosaccharides to suckling piglets affects intestinal microbiota colonization and immune development. J Anim Sci 2018;96:2139-53. [Crossref] [PubMed]
- Forster SC, Kumar N, Anonye BO, et al. A human gut bacterial genome and culture collection for improved metagenomic analyses. Nat Biotechnol 2019;37:186-92. [Crossref] [PubMed]
- Lagier JC, Khelaifia S, Alou MT, et al. Culture of previously uncultured members of the human gut microbiota by culturomics. Nat Microbiol 2016;1:16203. [Crossref] [PubMed]
- Lefterova MI, Suarez CJ, Banaei N, et al. Next-Generation Sequencing for Infectious Disease Diagnosis and Management: A Report of the Association for Molecular Pathology. J Mol Diagn 2015;17:623-34. [Crossref] [PubMed]
- Klindworth A, Pruesse E, Schweer T, et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 2013;41:e1. [Crossref] [PubMed]
- Laudadio I, Fulci V, Palone F, et al. Quantitative Assessment of Shotgun Metagenomics and 16S rDNA Amplicon Sequencing in the Study of Human Gut Microbiome. OMICS 2018;22:248-54. [Crossref] [PubMed]
- Morgan XC, Huttenhower C. Chapter 12: Human microbiome analysis. PLoS Comput Biol 2012;8:e1002808. [Crossref] [PubMed]
- Karlsson FH, Fåk F, Nookaew I, et al. Symptomatic atherosclerosis is associated with an altered gut metagenome. Nat Commun 2012;3:1245. [Crossref] [PubMed]
- Dinakaran V, Rathinavel A, Pushpanathan M, et al. Elevated levels of circulating DNA in cardiovascular disease patients: metagenomic profiling of microbiome in the circulation. PLoS One 2014;9:e105221. [Crossref] [PubMed]
- Fu J, Bonder MJ, Cenit MC, et al. The Gut Microbiome Contributes to a Substantial Proportion of the Variation in Blood Lipids. Circ Res 2015;117:817-24. [Crossref] [PubMed]
- Yin J, Liao SX, He Y, et al. Dysbiosis of Gut Microbiota With Reduced Trimethylamine-N-Oxide Level in Patients With Large-Artery Atherosclerotic Stroke or Transient Ischemic Attack. J Am Heart Assoc 2015;4:e002699. [Crossref] [PubMed]
- Feng Q, Liu Z, Zhong S, et al. Integrated metabolomics and metagenomics analysis of plasma and urine identified microbial metabolites associated with coronary heart disease. Sci Rep 2016;6:22525. [Crossref] [PubMed]
- Emoto T, Yamashita T, Sasaki N, et al. Analysis of Gut Microbiota in Coronary Artery Disease Patients: a Possible Link between Gut Microbiota and Coronary Artery Disease. J Atheroscler Thromb 2016;23:908-21. [Crossref] [PubMed]
- Kelly TN, Bazzano LA, Ajami NJ, et al. Gut Microbiome Associates With Lifetime Cardiovascular Disease Risk Profile Among Bogalusa Heart Study Participants. Circ Res 2016;119:956-64. [Crossref] [PubMed]
- Jie Z, Xia H, Zhong SL, et al. The gut microbiome in atherosclerotic cardiovascular disease. Nat Commun 2017;8:845. [Crossref] [PubMed]
- Li J, Zhao F, Wang Y, et al. Gut microbiota dysbiosis contributes to the development of hypertension. Microbiome 2017;5:14. [Crossref] [PubMed]
- Cui L, Zhao T, Hu H, et al. Association Study of Gut Flora in Coronary Heart Disease through High-Throughput Sequencing. Biomed Res Int 2017;2017:3796359.
- Menni C, Lin C, Cecelja M, et al. Gut microbial diversity is associated with lower arterial stiffness in women. Eur Heart J 2018;39:2390-7. [Crossref] [PubMed]
- Zhernakova DV, Le TH, Kurilshikov A, et al. Individual variations in cardiovascular-disease-related protein levels are driven by genetics and gut microbiome. Nat Genet 2018;50:1524-32. [Crossref] [PubMed]
- Yoshida N, Emoto T, Yamashita T, et al. Bacteroides vulgatus and Bacteroides dorei Reduce Gut Microbial Lipopolysaccharide Production and Inhibit Atherosclerosis. Circulation 2018;138:2486-98. [Crossref] [PubMed]
- Zhu Q, Gao R, Zhang Y, et al. Dysbiosis signatures of gut microbiota in coronary artery disease. Physiol Genomics 2018;50:893-903. [Crossref] [PubMed]
- Kurilshikov A, van den Munckhof ICL, Chen L, et al. Gut Microbial Associations to Plasma Metabolites Linked to Cardiovascular Phenotypes and Risk. Circ Res 2019;124:1808-20. [Crossref] [PubMed]
- Liu H, Chen X, Hu X, et al. Alterations in the gut microbiome and metabolism with coronary artery disease severity. Microbiome 2019;7:68. [Crossref] [PubMed]
- Rothschild D, Weissbrod O, Barkan E, et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 2018;555:210-5. [Crossref] [PubMed]
- Bennett BJ, de Aguiar Vallim TQ, Wang Z, et al. Trimethylamine-N-oxide, a metabolite associated with atherosclerosis, exhibits complex genetic and dietary regulation. Cell Metab 2013;17:49-60. [Crossref] [PubMed]
- Wang Z, Klipfell E, Bennett BJ, et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 2011;472:57-63. [Crossref] [PubMed]
- Wang Z, Roberts AB, Buffa JA, et al. Non-lethal Inhibition of Gut Microbial Trimethylamine Production for the Treatment of Atherosclerosis. Cell 2015;163:1585-95. [Crossref] [PubMed]
- Ma G, Pan B, Chen Y, et al. Trimethylamine N-oxide in atherogenesis: impairing endothelial self-repair capacity and enhancing monocyte adhesion. Biosci Rep 2017;37:BSR20160244. [Crossref] [PubMed]
- Makrecka-Kuka M, Volska K, Antone U, et al. Trimethylamine N-oxide impairs pyruvate and fatty acid oxidation in cardiac mitochondria. Toxicol Lett 2017;267:32-8. [Crossref] [PubMed]
- Li XS, Wang Z, Cajka T, et al. Untargeted metabolomics identifies trimethyllysine, a TMAO-producing nutrient precursor, as a predictor of incident cardiovascular disease risk. JCI Insight 2018;3:e99096. [Crossref] [PubMed]
- Koeth RA, Levison BS, Culley MK, et al. γ-Butyrobetaine is a proatherogenic intermediate in gut microbial metabolism of L-carnitine to TMAO. Cell Metab 2014;20:799-812. [Crossref] [PubMed]
- Ríos-Covián D, Ruas-Madiedo P, Margolles A, et al. Intestinal Short Chain Fatty Acids and their Link with Diet and Human Health. Front Microbiol 2016;7:185. [Crossref] [PubMed]
- Koh A, De Vadder F, Kovatcheva-Datchary P, et al. From Dietary Fiber to Host Physiology: Short-Chain Fatty Acids as Key Bacterial Metabolites. Cell 2016;165:1332-45. [Crossref] [PubMed]
- Ohira H, Tsutsui W, Fujioka Y. Are Short Chain Fatty Acids in Gut Microbiota Defensive Players for Inflammation and Atherosclerosis? J Atheroscler Thromb 2017;24:660-72. [Crossref] [PubMed]
- Pluznick JL. Renal and cardiovascular sensory receptors and blood pressure regulation. Am J Physiol Renal Physiol 2013;305:F439-F444. [Crossref] [PubMed]
- Aguilar EC, Leonel AJ, Teixeira LG, et al. Butyrate impairs atherogenesis by reducing plaque inflammation and vulnerability and decreasing NFκB activation. Nutr Metab Cardiovasc Dis 2014;24:606-13. [Crossref] [PubMed]
- Moris D, Giaginis C, Tsourouflis G, et al. Farnesoid-X Receptor (FXR) as a Promising Pharmaceutical Target in Atherosclerosis. Curr Med Chem 2017;24:1147-57. [Crossref] [PubMed]
- Kawamata Y, Fujii R, Hosoya M, et al. A G protein-coupled receptor responsive to bile acids. J Biol Chem 2003;278:9435-40. [Crossref] [PubMed]
- de Boer JF, Schonewille M, Boesjes M, et al. Intestinal Farnesoid X Receptor Controls Transintestinal Cholesterol Excretion in Mice. Gastroenterology 2017;152:1126-1138.e6. [Crossref] [PubMed]
- Westin S, Heyman RA, Martin R. FXR, a therapeutic target for bile acid and lipid disorders. Mini Rev Med Chem 2005;5:719-27. [Crossref] [PubMed]
- Miyazaki-Anzai S, Masuda M, Kohno S, et al. Simultaneous inhibition of FXR and TGR5 exacerbates atherosclerotic formation. J Lipid Res 2018;59:1709-13. [Crossref] [PubMed]
- Koren O, Spor A, Felin J, et al. Human oral, gut, and plaque microbiota in patients with atherosclerosis. Proc Natl Acad Sci U S A 2011;108 Suppl 1:4592-8. [Crossref] [PubMed]
- Chacón MR, Lozano-Bartolomé J, Portero-Otín M, et al. The gut mycobiome composition is linked to carotid atherosclerosis. Benef Microbes 2018;9:185-98. [Crossref] [PubMed]
- Lam V, Su J, Hsu A, et al. Intestinal Microbial Metabolites Are Linked to Severity of Myocardial Infarction in Rats. PLoS One 2016;11:e0160840. [Crossref] [PubMed]
- Ohashi R, Mu H, Wang X, et al. Reverse cholesterol transport and cholesterol efflux in atherosclerosis. QJM 2005;98:845-56. [Crossref] [PubMed]
- Lan TH, Huang XQ, Tan HM. Vascular fibrosis in atherosclerosis. Cardiovasc Pathol 2013;22:401-7. [Crossref] [PubMed]
- Chambers ES, Preston T, Frost G, et al. Role of Gut Microbiota-Generated Short-Chain Fatty Acids in Metabolic and Cardiovascular Health. Curr Nutr Rep 2018;7:198-206. [Crossref] [PubMed]
- Zhou X, Li J, Guo J, et al. Gut-dependent microbial translocation induces inflammation and cardiovascular events after ST-elevation myocardial infarction. Microbiome 2018;6:66. [Crossref] [PubMed]
- Li XS, Obeid S, Wang Z, et al. Trimethyllysine, a trimethylamine N-oxide precursor, provides near- and long-term prognostic value in patients presenting with acute coronary syndromes. Eur Heart J 2019;40:2700-9. [Crossref] [PubMed]
- Amar S, Engelke M. Periodontal innate immune mechanisms relevant to atherosclerosis. Mol Oral Microbiol 2015;30:171-85. [Crossref] [PubMed]
- Ott SJ, El Mokhtari NE, Musfeldt M, et al. Detection of diverse bacterial signatures in atherosclerotic lesions of patients with coronary heart disease. Circulation 2006;113:929-37. [Crossref] [PubMed]
- Renvert S, Pettersson T, Ohlsson O, et al. Bacterial profile and burden of periodontal infection in subjects with a diagnosis of acute coronary syndrome. J Periodontol 2006;77:1110-9. [Crossref] [PubMed]
- Zeng XT, Leng WD, Lam YY, et al. Periodontal disease and carotid atherosclerosis: A meta-analysis of 17,330 participants. Int J Cardiol 2016;203:1044-51. [Crossref] [PubMed]
- Shi Q, Zhang B, Huo N, et al. Association between Myocardial Infarction and Periodontitis: A Meta-Analysis of Case-Control Studies. Front Physiol 2016;7:519. [Crossref] [PubMed]
- Lam V, Su J, Koprowski S, et al. Intestinal microbiota determine severity of myocardial infarction in rats. FASEB J 2012;26:1727-35. [Crossref] [PubMed]
- Chen L, Liu W, Li Y, et al. Lactobacillus acidophilus ATCC 4356 attenuates the atherosclerotic progression through modulation of oxidative stress and inflammatory process. Int Immunopharmacol 2013;17:108-15. [Crossref] [PubMed]
- Li J, Lin S, Vanhoutte PM, et al. Akkermansia Muciniphila Protects Against Atherosclerosis by Preventing Metabolic Endotoxemia-Induced Inflammation in Apoe-/- Mice. Circulation 2016;133:2434-46. [Crossref] [PubMed]
- Qiu L, Tao X, Xiong H, et al. Lactobacillus plantarum ZDY04 exhibits a strain-specific property of lowering TMAO via the modulation of gut microbiota in mice. Food Funct 2018;9:4299-309. [Crossref] [PubMed]
- Wang D, Xia M, Yan X, et al. Gut microbiota metabolism of anthocyanin promotes reverse cholesterol transport in mice via repressing miRNA-10b. Circ Res 2012;111:967-81. [Crossref] [PubMed]
- Matziouridou C, Marungruang N, Nguyen TD, et al. Lingonberries reduce atherosclerosis in Apoe(-/-) mice in association with altered gut microbiota composition and improved lipid profile. Mol Nutr Food Res 2016;60:1150-60. [Crossref] [PubMed]
- Chen ML, Yi L, Zhang Y, et al. Resveratrol Attenuates Trimethylamine-N-Oxide (TMAO)-Induced Atherosclerosis by Regulating TMAO Synthesis and Bile Acid Metabolism via Remodeling of the Gut Microbiota. mBio 2016;7:e02210-15. [Crossref] [PubMed]
- Zhu L, Zhang D, Zhu H, et al. Berberine treatment increases Akkermansia in the gut and improves high-fat diet-induced atherosclerosis in Apoe-/- mice. Atherosclerosis 2018;268:117-26. [Crossref] [PubMed]
- Cammarota G, Ianiro G, Gasbarrini A. Fecal microbiota transplantation for the treatment of Clostridium difficile infection: a systematic review. J Clin Gastroenterol 2014;48:693-702. [Crossref] [PubMed]
- Hu XF, Zhang WY, Wen Q, et al. Fecal microbiota transplantation alleviates myocardial damage in myocarditis by restoring the microbiota composition. Pharmacol Res 2019;139:412-21. [Crossref] [PubMed]
- Trøseid M. Gut microbiota and acute coronary syndromes: ready for use in the emergency room? Eur Heart J 2017;38:825-7. [Crossref] [PubMed]
- Hansson GK, Hermansson A. The immune system in atherosclerosis. Nat Immunol 2011;12:204-12. [Crossref] [PubMed]


